| A Phylogeny of Cichlids |
| A Phylogeny of Cichlids |
A phylogeny is an expression of the evolutionary relationships of a group of animals. The purpose of a phylogeny is to illustrate which taxa (e.g., species, genera, etc) are most closely related to which other ones, and also, to illustrate the evidence that we have to support that relationship. The science of discovering those relationships is called systematics and a scientist who does this kind of work is a systematist.
An important point to remember is that there is a SINGLE, TRUE phylogeny, i.e., there was a particular path which evolution took leading to the current diversity of organisms inhabiting the earth. The problem comes in trying to reconstruct that pattern.
As scientists gather more data and reanalyze older data in light of new ideas, our understanding of these relationships grows. We might realize that creatures which we once thought were closely related are not in fact so closely related at all. Thus the "current" phylogeny may change, sometimes dramatically. [Click here to see a current summary of the entire TREE OF LIFE].
Some scientists are interested in discovering the true phylogeny of groups of organisms, such as the cichlids. We can then use that information to understand the evolution of particular characters. For example, with a good understanding of cichlid phylogeny, we can see that mouthbrooding has evolved multiple times in cichlids, not just once as you might think.
But, other people that work with, keep, buy or sell cichlids have a more utilitarian need for arranging fish. They need a set of names to apply to the fish so that everyone knows which fish is being discussed. We call this set of names a classification. The science of applying names to organisms is called taxonomy. Ideally, the classification should reflect the phylogeny (although not everyone agrees with this ideal). The problem comes when we try to create a classification with our imperfect knowledge of phylogeny. The result is that as our knowledge of phylogeny changes, so too must the classification, and therefore, so too must the names by which we call these fish.
The current classification of cichlids (and indeed of all organisms) is not perfect. In fact, it is a hybrid between older ideas about how organisms were related and newer ideas. Many scientists would dearly like to scrap the old system of Kingdom, Phylum, Class, Order, Family, Genus and Species, because that system does not accurately portray our current understanding of evolutionary relationships. However, there is no widely accepted alternative available to take its place. So in the meantime, we continue to tinker with the old system trying to make it at least partially represent our knowledge.
Click to see the current classification of fishes, and of cichlids.
Methods of reconstructing a phylogeny
Since we were not present when most cichlids evolved, we cannot know for sure which came from which. Indeed even if we were present, this would be a difficult task given our tendency to stay above-water most of the time. The fossil record is also of very limited use for discovering cichlid ancestry so instead we must rely on an indirect method of determining which species is most closely related to which other species.
The method of reconstructing phylogenies we use is called phylogenetic systematics, or often abbreviated to cladistics. Cladistics is based on the principle of shared derived characters, often called synapomorphies. If two organisms share a very peculiar trait, we assume that the reason they have this trait is because their common ancestor had it. For example, imagine two fish species that both have a bright red fluorescent stripe along the side. The odds that such a peculiar trait evolved twice is very low. It is far more likely that the common ancestor to these two fishes had the bright red fluorescent stripe and passed it on to its offspring. So we call this bright red fluorescent stripe a shared derived character for these two species.
Now imagine that we are examining four fish species. Two species share the peculiar trait of having extremely long pelvic fins (the fins on the underside of the body), three have the bright red flourescent stripe and the fourth has neither of these peculiar traits. All four species have a pointy tail.
A cladist (a person who analyzes phylogenies using cladistics) would hypothesize that the two with the extremely long pelvic fins are closest relatives (sister taxa). The third species, which does not have long pelvic fins, but shares the bright red fluroescent stripe with the other two, is the sister group to the group consisting of the first two species. We know all four species form a group because they all share the derived character of having a pointy tail.
We can illustrate the data as follows:
Species
A B C D
trait
long pelvics yes yes no no
red stripe yes yes yes no
pointy tail yes yes yes yes
The key to doing cladistics is identifying shared derived characters. Shared common characters, such as having two eyes are of no value to a cladist because lots of organisms have two eyes. Having two eyes tells us nothing about relationships.
A shorthand way to represent this tree is with the notation (((A B) C) D). This notation tells us we have a tree with A and B as sister groups, and together they form the sister group to C, and together the group (A B) C forms the sister group to D.
What kind of characters are used in cladistics?
Traditionally, the kinds of things used in a cladistic analysis were what are often called hard parts, i.e. things like bones, scales and teeth. More recently, soft tissues like muscles have proved very useful. Very recently (the last ten years) a number of researchers have started using molecular data, e.g. portions of the DNA sequence extracted from the cells of cichlids. Despite what many say (and argue), I feel that no one source of data is the ultimate solution. For example, many people put immense hope that once we know the DNA sequence of a an organism, we know everything we need to know. This isn't true. There are many troubling problems with interpreting sequence data that are often overlooked by newcomers to the field, but weigh heavily on the minds of many cladists. For the latest arguments about this important issue, find a copy of the latest issue of the journal Cladistics in your nearest university library.
Reading a phylogeny
The important point when reading a phylogeny is to see which taxa branch from the same common point (called a node). If two taxa branch from the same node, they are called sister taxa, meaning they are closest relatives. If more than two taxa branch from the same node, that means we currently have insufficient data to determine which of those taxa are more closely related to each other than the others. We may call that node unresolved.
The words monophyletic group are important words that you often see in reference to phylogenies. In technical terms, a monophyletic group means a group consisting of a node, plus all its descendents. In practical terms you can think of a phylogeny as a tree. If you grab the tree at any point and break off a chunk and hold it in your hand, you are holding a monophyletic group. If you break off another piece from the main tree and combine it with your first piece, you no longer have a monophyletic group. Similarly, if you take a piece out of your original chunk and throw it away, the original chunk is no longer a monophyletic group, though the little piece you threw away was a monophyletic group. A monophyletic group includes ALL the descendents of a node, no more and no less.
Imagine that there was a small ancestral cichlid living in South America from which evolved all the current Apistogramma species. A group consisting of the node plus all its descendents, including all the current Apistogramma species, would be considered monophyletic. But what if recently one of the Apistogramma species evolved to be quite large and no longer even looked like a typical Apistogramma. We might be tempted to exclude it from the group. To do so would be improper from the point of view of understanding relationships. In doing so, we would have made our group paraphyletic, i.e. not including all the descendents.
To return to our analogy of grabbing a branch of a tree, if you understand what I have written so far, you will appreciate that a monophyletic group can (and does) contain within itself many monophyletic groups.
The goal of modern cladistics is to identify monophyletic groups.
Cichlids have proved particularly troublesome for reconstructing phylogeny. Much evolution in various lines of cichlids has occurred in short periods of time, making the job of figuring out which species came from which others rather difficult.
Here, I present Melanie Stiassny's 1991 phylogeny of the major groups of cichlids. Notice the unresolved node joining the bulk of the cichlids. There is also much debate about the relationships within each of these groups, e.g. the relationships within the Neotropical cichlids, and even if the Neotropical cichlids are in fact a monophyletic group.
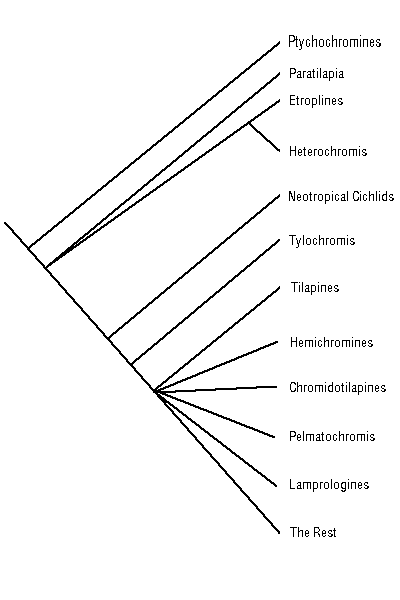
Modified From: Stiassny, M.L.J. (1991) Phylogenetic intrarelationships of the family Cichlidae: an overview. Pp. 1-35 in: Cichld Fishes: Behaviour, ecology and evolution (M.H.A. Keenleyside, Ed), Chapman and Hall, New York.
Further Information
For more information on phylogenetic systematics, how it is done and what it means, try the UC Berkeley Museum of Paleontology "Journey into Phylogenetic Systematics"
![[Go Back to Contents]](gifs/back.gif) |
![[Send email to Ron Coleman]](gifs/address.gif) |